
Knockdown of carboxypeptidase A4 (CPA4) inhibits gastric cancer cell progression via cell cycle arrest and apoptosis
Introduction
Gastric cancer is one of the most prevalent cancers, with more than one million cases diagnosed worldwide annually (1). It was reported that males are more susceptible to developing gastric cancer from birth to age 74 compared with females (1). Gastric cancer accounts for 8.3% of all cancer-related deaths (2). The 5-year survival rate was shown to vary in patients with different stages during surgical intervention, with 94% at stage IA, 88% at stage IB, and only 18% at stage IIIC (3). Therefore, it is of great importance to identify key biomarkers for evaluating gastric cancer progression, as well as exploring potential therapeutic targets for gastric cancer therapy.
Carboxypeptidase A4 (CPA4) is a member of the Zn-containing metallocarboxypeptidase (MCP) family. It catalyzes the release of carboxy-terminal amino acids. It has been confirmed that CPA4, an important regulator of inflammation, plays an important role in tumor microenvironment (TME) formation and distant metastasis in tumor (4,5). CPA4 has been shown to be upregulated in several cancers, and is speculated to be involved in regulating the progression of several cancers, including non-small cell lung cancer (6), pancreatic cancer (7), bladder cancer (8), colorectal cancer (9), hepatocellular carcinoma (10), and breast cancer (11). For example, CPA4 expression was associated with poor prognoses by promoting cell proliferation in hepatocellular carcinoma (10). Upregulated CPA4 expression was an independent prognostic factor for patients with liver cancer (12). Sun et al. (13) demonstrated that CPA4 is associated with poor prognosis in patients with gastric cancer. However, the regulatory role of CPA4 in gastric cancer is still unknown.
In the present study, we explored the effects of CPA4 on gastric cancer progression in vitro. This study may provide a foundation for further study into therapeutic targets for gastric cancer. We present the following article in accordance with the MDAR reporting checklist (available at https://jgo.amegroups.com/article/view/10.21037/jgo-22-987/rc).
Methods
Cell culture
Human gastric adenocarcinoma cell line (AGS cell line) were obtained from the American Type Culture Collection (ATCC; Rockville, MD, USA) and cultured in RPMI-1640 culture medium (Thermo Fisher Scientific, Waltham, MA, USA) at 37 ℃ with 5% CO2.
Construction of lentiviruses for CPA4 knockdown
The target insert sequence of CPA4 (5'-TTCTCCGAACGTGTCACGT-3') was designed by Shanghai GeneChem Co. Ltd. Two single-chain DNA sequences were designed by Shanghai Generay Biotech Co., Ltd.: shCPA4S1, 5'-CCGGGCGTATGACAACGGCATCAAATTCAAGAGATTTGATGCCGTTGTCATACGCTTTTTG-3'; and shCPA4S2, 5'-AATTCAAAAAGCGTATGACAACGGCATCAAATCTCTTGAATTTGATGCCGTTGTCATACGC-3'. Single-chain DNA was dissolved into annealing buffer (20 µM) and incubated in a water bath at 90 ℃ for 15 min. After being naturally cooled to room temperature, a double strand with sticky ends was formed. The GV115 lentiviral vector was linearized by AgeI (#R3552L; New England BioLabs, USA) and EcoRI (#R3101L; New England BioLabs, USA). Then, the DNA sequences were ligated into the linearized GV115 lentiviral vector, and the resulting plasmid was used to transform top 10 E. coli competent cells (#CB104-03; TIANGEN, Beijing, China). The plasmid was extracted from the top 10 E. coli competent cells and verified by restriction endonuclease digestion followed by Sanger sequencing (Genscript Co., Ltd.; Nanjing, China). Transfection was performed using Lipofectamine® 2000 (Invitrogen; Thermo Fisher Scientific, Inc., USA). We collected the supernatants containing lentiviruses after 48 h of culture and purified them at 1,000 ×g and 4 ℃ for 2 min (Himac CT15RE; Hitachi).
Quantitative real-time PCR (qRT-PCR) analysis
The expression level of CPA4 was evaluated in AGS and transfected cells. The total RNA in transfected cells was extracted by TRIzol (Invitrogen, Carlsbad, CA, USA). The primers were designed and synthesized by Sangon (Shanghai, China). The primers are listed as follows: CPA4 forward primer: 5'-ATTGGACATTCGTTTGAAAACCG-3'; CPA4 reverse primer: 5'-GGGAGATCCACTCTCGGGA-3'. GAPDH forward primer: 5'- TGACTTCAACAGCGACACCCA-3'; GAPDH reverse primer: 5'- CACCCTGTTGCTGTAGCCAAA-3'. Then, qRT-PCR analysis was performed by applying the HiScript II One Step qRT-PCR SYBR Green Kit (#Q221-01), tested on an ABI 7900 system (Foster City, CA, USA). Expression was normalized to GAPDH and calculated using the 2–ΔCt method.
Western blot analysis
The proteins were isolated by RIPA lysis buffer (#R0278, Sigma). The standard sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) method was applied to separate the proteins (14). Then, the gels were transferred, blocked, and cultured with primary antibodies against the following: CPA4 (1:300; #ab81543; Abcam); epithelial-mesenchymal transition (EMT)-related transcription factors NFκB p65 (1:2,000; #8242; CST), p-NFκB p65 (1:1,000; #3033; CST), Slug (1:1,000; #9585; CST), Fibronectin (1:500; MAB1918; R&D), N-cadherin (1:1,000; ab18203; Abcam), Vimentin (1:1,000; #3932; CST), and E-cadherin (1:1,000; #14472; CST); multiple malignancy-associated genes Twist (1:100; ab50887; Abcam), c-Myc (1:1,000; ab32072; Abcam), and Snail (1:2,000; #3879; CST); Wnt signaling pathway proteins β-catenin (1:2,000; #8480; CST) and p-β-catenin (1:1,000; #2009; CST); PI3K/AKT/mTOR signaling pathway proteins AKT (1:2,000; ab179463; Abcam), p-AKT (1:1,000; #4060; CST), mTOR (1:500; #2972; CST), and p-mTOR (1:2,000; ab109268; Abcam); MAPK signaling pathway proteins p38 (1:2,000; #8690; CST) and p-p38 (1:1,000; #4631; CST); and tumor migration and invasion-related proteins ERK (1:2,000; ab17942; Abcam), p-ERK (1:2,000; #4370; CST), MMP-2 (1:500; #40994; CST), and MMP-9 (1:500; #13667; CST). GAPDH (1:10,000; #sc-32233; Santa-Cruz) was used as the internal control. The membranes were washed. Then they were cultured with HRP-goat anti rabbit secondary antibody (1:10,000; AS1107; ASPEN) and HRP-goat anti mouse (1:10,000; AS1106; ASPEN). Finally, the bands were observed under ECL plus and X-ray film.
Protein-protein interaction (PPI) network
According to the significantly dysregulated proteins in shCPA4 compared with shCtrl cells, a PPI network of CPA4 was generated. Analysis of the interaction network was performed using the STRING database (https://string-db.org/).
Cell colony formation analysis
AGS cells in logarithmic growth phase were subjected to cell trypsin digestion (#25200-720; Gibco), resuspended, then counted. About 500 cells/well were seeded in a 6-well plate, and each sample was performed in triplicate. After culture for 14 days, the cells were washed with phosphate-buffered saline (PBS), fixed with 4% paraformaldehyde for 30–60 min, and washed with PBS again. Then, the cells were stained with impurity-free crystal violet staining solution for 10–20 min, washed with ddH2O, and photographed under a fluorescence microscope (XDS-100; Caikon, Shanghai, China).
Cell proliferation assay
AGS cells in logarithmic growth phase were subjected to cell trypsin digestion (#25200-720; Gibco; Thermo Fisher Scientific, Inc., USA), resuspended, then counted. Approximately 2,000 cells/well were seeded in each well of a 6-well plate and cultured at 37 ℃ with 5% CO2. Celigo® Image Cytometer (Nexcelom) was used to evaluate the number of cells by scanning green fluorescence daily for 5 days at room temperature (15).
Cell migration and invasion analyses
The Transwell assays were applied to evaluate the cell migration and invasion of cells. A 24-well Transwell chamber and 8.0-µm pore membranes (Corning, USA) coated with Matrigel (BD Biosciences) were used for the cell invasion assay. Approximately 1.0×105 cells/well in 100 mL serum-free Dulbecco’s modified Eagle’s medium (DMEM) were seeded in the top chamber for 6 h at 37 ℃. Then 500 mL medium containing 30% fetal bovine serum (FBS) was placed into the lower chamber. The Transwell was washed, fixed with 4% paraformaldehyde, and stained with 0.1% crystal violet. Eventually, 5 visual fields were randomly selected, and the cell numbers were counted under a 400× microscope. The cell migration assay was similar to that of invasion assay except that the chamber was not coated with Matrigel.
Cell apoptosis analysis
About 5.0×105 cells were added into a 6-well plate and maintained at 37 ℃ with 5% CO2 until the cell growth coverage reached 70%. The cells were collected, washed with pre-chilled D-Hanks, and treated with 1× binding buffer. Then, the cells were stained with Annexin V-APC for 10–15 min and observed under a fluorescence microscope (IX71, OLYMPUS, Tokyo, Japan).
Cell cycle assay
About 2.0×106 cells were seeded into a 6-cm dish after lentiviral infection for 5 days. The dishes were maintained at 37 ℃ for 40 h until the confluence of cells reached about 80%. The cells were washed with pre-chilled D-Hanks (pH 7.2–7.4) and fixed with 70% ethanol overnight at 4 ℃. After digestion with RNaseA, the cells were labeled with a solution of 40× PI, 100× RNase, and 1× D-Hanks at a ratio of 25:10:1,000 and were analyzed with Cell Lab Quanta Beckman Coulter to measure fluorescence. The cell cycle was analyzed by ModFit LT 3.2 (Verity Software House).
Statistical analysis
GraphPad Prism 8 (San Diego, CA, USA) was used for the statistical analysis and graph drawing. Student’s t-test or one-way ANOVA was performed for comparative analyses depending on the number of groups. Data are exhibited as mean ± standard deviation (SD). P<0.05 indicated that the difference was statistically significant. *, P<0.05; **, P<0.01; ***, P<0.001.
Results
CPA4 is highly expressed in gastric cancer cell lines
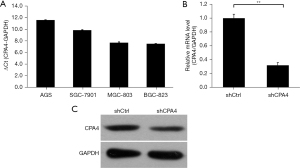
The expression of CPA4 was significantly upregulated in the gastric cancer cell lines AGS, SGC-7901, MGC-803, and BGC-823, with the highest expression level in AGS cells and the lowest in MGC-803 cells (Figure 1A). After lentivirus transfection with shCPA4, the mRNA and protein expression levels of CPA4 were significantly decreased (P<0.01; Figure 1B,1C), which demonstrated the successful construction of knockdown samples.
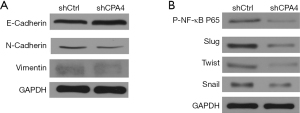
Overexpressed CPA4 is associated with induction of EMT
The role of CPA4 in gastric cancer was assessed by testing the expression levels of EMT-related markers in transfected cells. In the western blot analysis, we found that E-cadherin was upregulated, while N-cadherin and Vimentin were downregulated in shCPA4 cells (Figure 2A), indicating that overexpressed CPA4 was associated with the induction of EMT. As transcription factors of EMT, p-NFκB-p65, Slug, Twist, and Snail protein expression levels were significantly downregulated in shCPA4 compared with shCtrl cells (Figure 2B). These results demonstrated that CPA4 promotes EMT processes in gastric cancer cells.
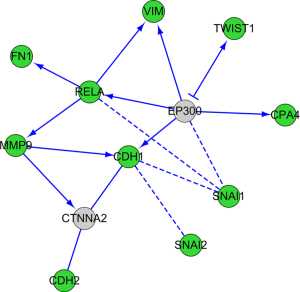
PPI network analysis
The regulatory relationship among TWIST1, FN1, SNAI1, SNAI2, VIM, MMP9, RELA, CDH1, CDH2, and CPA4 was carried using STRING database (Figure 3). CPA4 was activated by EP300, and it also activates VIM, RELA, SNAI1 and CDH1. The results provide insights into the potential regulatory relationship of CPA4 in gastric cancer.
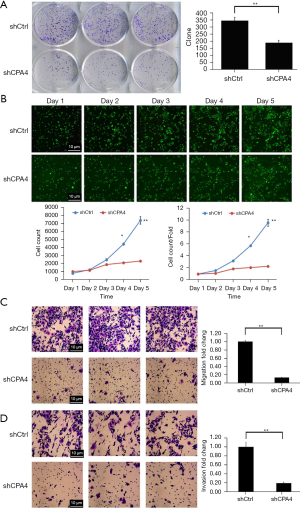
Knockdown of CPA4 inhibits cell colony formation, proliferation, migration, and invasion of gastric cancer cells
The colonies of shCPA4 cells were significantly lower than those of shCtrl cells (P<0.01; Figure 4A). The cell count and cell count/fold ratio in the shCPA4 group were significantly lower than those in shCtrl cells (P<0.01; Figure 4B). The results indicated that CPA4 promoted the proliferation of gastric cancer cells. The numbers of migrating and invading cells were significantly repressed by shCPA4 (P<0.01; Figure 4C,4D). Overall, the results demonstrate that knockdown of CPA4 inhibits cell colony formation, proliferation, migration, and invasion of gastric cancer cells.
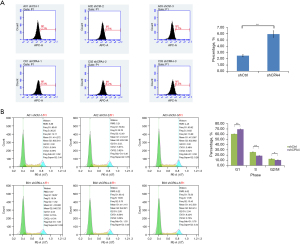
Knockdown of CPA4 promotes cell apoptosis in gastric cancer
ShCPA4 significantly increased the number of apoptotic cells compared with shCtrl (P<0.01; Figure 5A), demonstrating that shCPA4 promotes cell apoptosis. Compared with shCtrl, shCPA4 significantly decreased the cells in S (25.88% vs. 18.66%; P<0.05) and G2/M phase (12.83% vs. 10.57%; P<0.05), and increased those in G1 phase (60.39% vs. 69.43%; P<0.05; Figure 5B). These results demonstrated that knockdown of CPA4 promoted the cell apoptosis of gastric cancer cells due to its arrest of cell cycle in G1 phase.
Discussion
Gastric cancer is a common cancer with a high survival rate at the early stage. However, the 5-year overall survival rate significantly decreases when patients are diagnosed at the later stage. Therefore, exploring useful biomarkers for gastric cancer progression and therapeutic targets are of great importance. In the present study, we investigated the role of CPA4 in gastric cancer progression in vitro. We found that: (I) CPA4 was significantly upregulated in gastric cancer cells; (II) knockdown of CPA4 inhibits cell proliferation, and cell migration and invasion, and promotes cell apoptosis and arrests the cell cycle at G1 phase; (III) CPA4 regulates gastric cancer progression through EMT processes.
In the present study, CPA4 was upregulated in gastric cancer cells, which was consistent with a report by Sun et al. (13). They analyzed the relationship between overexpressed CPA4 and clinical factors such as tumor size, stage, lymph node metastasis, depth of invasion, and distant metastasis and demonstrated that overexpressed CPA4 can be utilized as an independent poor prognostic factor in gastric cancer. In our present study, we lacked clinical verification, but emphasized the role of CPA4 in gastric cancer progression. We demonstrated that overexpressed CPA4 promotes cancer cell progression. In other cancers, CPA4 was also reported to be upregulated, and overexpressed CPA4 has been shown to be involved in cancer cell progression. Zhang et al. (10) revealed that CPA4 promotes the proliferation and stem cell characteristics of hepatocellular carcinoma. Downregulation of CPA4 inhibits the cell growth of non-small cell lung cancer through the AKT/c-MYC pathway (16). Shao et al. (7) demonstrated that overexpressed CPA4 promotes pancreatic cancer via stimulating PI3K/AKT/mTOR signaling. Overall, we can conclude that CPA4 exhibited similar functions in gastric cancer cells as in other cancers.
As mentioned above, CPA4 regulates cancer through various signaling pathways. However, in the present study, we also found that EMT processes were induced. EMT is an important process for vertebrate embryonic development, driven by transcription factors such as SNAIL1, TWIST1, and ZEB1 in mesenchymal programs during embryogenesis, and deletion of SNAIL1 in mice results in embryonic lethality with impaired mesoderm development (17,18). In the present study, the expression levels of SNAIL and TWIST were significantly downregulated by shCPA4. The EMT-related biomarkers E-cadherin, N-cadherin, and Vimentin were dysregulated by shCPA4. These results demonstrated that CPA4 induces EMT in gastric cancer cells. EMT is involved in the progression of many cancers. It may also be involved in gastric cancer progression by arresting the cell cycle at G1 phase. Lovisa et al. (19) found that transgenic expression of either TWIST or SNAIL is sufficient to promote prolonged TGF-β1-induced G2 arrest of tubular epithelial cells, limiting the cells’ potential for repair and regeneration. SNAIL can block the cell cycle and confers resistance to cell death (20). Yang et al. (21) demonstrated that the role of cell cycle state in the regulation of TGF-β-induced EMT and apoptosis, and also confirmed that TGF-β induced EMT via a cell cycle-dependent mechanism. Therefore, we can assume that EMT is regulated by cell cycle arrest caused by the dysregulation of CPA4. Altogether, CPA4 was upregulated in gastric cancer cells, and overexpressed CPA4 promotes cancer cell progression through EMT by inducing cell cycle arrest. We speculate that administering a molecule that can specifically inhibit the transcription or translation of the CPA4 gene in tumor cells, or specifically inhibit the expression or activity of the CPA4 protein, may inhibit the growth, proliferation, differentiation, and/or survival of tumor cells. However, the regulation process is a complex scientific issue that requires more experiments for verification, as in vivo experiments are lacking.
Conclusions
The present study confirmed the role and function of CPA4 in gastric cancer in vitro. It has revealed that CPA4 was significantly upregulated in AGS gastric cancer cells, and knockdown of CPA4 was related to inhibited cell progression via arresting the cell cycle and inducing EMT.
Acknowledgments
Funding: This study was supported by the Medical Scientific Research Foundation of Zhejiang Province (No. 2021KY594); and Beijing Bethune Charitable Foundation of China (No. WCJZL202111).
Footnote
Reporting Checklist: The authors have completed the MDAR reporting checklist. Available at https://jgo.amegroups.com/article/view/10.21037/jgo-22-987/rc
Data Sharing Statement: Available at https://jgo.amegroups.com/article/view/10.21037/jgo-22-987/dss
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at https://jgo.amegroups.com/article/view/10.21037/jgo-22-987/coif). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Rawla P, Barsouk A. Epidemiology of gastric cancer: global trends, risk factors and prevention. Prz Gastroenterol 2019;14:26-38. [Crossref] [PubMed]
- Bray F, Ferlay J, Soerjomataram I, et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 2018;68:394-424. [Crossref] [PubMed]
- Howlader N, Noone A, Krapcho M, et al. SEER cancer statistics review, 1975–2012. National Cancer Institute. 2015.
- Ross PL, Cheng I, Liu X, et al. Carboxypeptidase 4 gene variants and early-onset intermediate-to-high risk prostate cancer. BMC Cancer 2009;9:69. [Crossref] [PubMed]
- Kayashima T, Yamasaki K, Yamada T, et al. The novel imprinted carboxypeptidase A4 gene (CPA4) in the 7q32 imprinting domain. Hum Genet 2003;112:220-6. [Crossref] [PubMed]
- Yan P, Lyu X, Wang S, et al. Insufficient ablation promotes the metastasis of residual non-small cell lung cancer (NSCLC) cells via upregulating carboxypeptidase A4. Int J Hyperthermia 2021;38:1037-51. [Crossref] [PubMed]
- Shao Q, Zhang Z, Cao R, et al. CPA4 Promotes EMT in Pancreatic Cancer via Stimulating PI3K-AKT-mTOR Signaling. Onco Targets Ther 2020;13:8567-80. [Crossref] [PubMed]
- Wei C, Zhou Y, Xiong Q, et al. Comprehensive Analysis of CPA4 as a Poor Prognostic Biomarker Correlated with Immune Cells Infiltration in Bladder Cancer. Biology (Basel) 2021;10:1143. [Crossref] [PubMed]
- Pan H, Pan J, Ji L, et al. Carboxypeptidase A4 promotes cell growth via activating STAT3 and ERK signaling pathways and predicts a poor prognosis in colorectal cancer. Int J Biol Macromol 2019;138:125-34. [Crossref] [PubMed]
- Zhang H, Hao C, Wang H, et al. Carboxypeptidase A4 promotes proliferation and stem cell characteristics of hepatocellular carcinoma. Int J Exp Pathol 2019;100:133-8. [Crossref] [PubMed]
- Wang Y, Xie Y, Niu Y, et al. Carboxypeptidase A4 negatively correlates with p53 expression and regulates the stemness of breast cancer cells. Int J Med Sci 2021;18:1753-9. [Crossref] [PubMed]
- Sun L, Guo C, Burnett J, et al. Association between expression of Carboxypeptidase 4 and stem cell markers and their clinical significance in liver cancer development. J Cancer 2017;8:111-6. [Crossref] [PubMed]
- Sun L, Guo C, Yuan H, et al. Overexpression of carboxypeptidase A4 (CPA4) is associated with poor prognosis in patients with gastric cancer. Am J Transl Res 2016;8:5071-5. [PubMed]
- Li XM, Yu WY, Chen Q, et al. LncRNA TUG1 exhibits pro-fibrosis activity in hypertrophic scar through TAK1/YAP/TAZ pathway via miR-27b-3p. Mol Cell Biochem 2021;476:3009-20. [Crossref] [PubMed]
- Nabzdyk CS, Chun M, Pradhan L, et al. High throughput RNAi assay optimization using adherent cell cytometry. J Transl Med 2011;9:48. [Crossref] [PubMed]
- Fu Y, Su L, Cai M, et al. Downregulation of CPA4 inhibits non small-cell lung cancer growth by suppressing the AKT/c-MYC pathway. Mol Carcinog 2019;58:2026-39. [Crossref] [PubMed]
- Nieto MA. Context-specific roles of EMT programmes in cancer cell dissemination. Nat Cell Biol 2017;19:416-8. [Crossref] [PubMed]
- Imani S, Hosseinifard H, Cheng J, et al. Prognostic Value of EMT-inducing Transcription Factors (EMT-TFs) in Metastatic Breast Cancer: A Systematic Review and Meta-analysis. Sci Rep 2016;6:28587. [Crossref] [PubMed]
- Lovisa S, LeBleu VS, Tampe B, et al. Epithelial-to-mesenchymal transition induces cell cycle arrest and parenchymal damage in renal fibrosis. Nat Med 2015;21:998-1009. [Crossref] [PubMed]
- Vega S, Morales AV, Ocaña OH, et al. Snail blocks the cell cycle and confers resistance to cell death. Genes Dev 2004;18:1131-43. [Crossref] [PubMed]
- Yang Y, Pan X, Lei W, et al. Transforming growth factor-beta1 induces epithelial-to-mesenchymal transition and apoptosis via a cell cycle-dependent mechanism. Oncogene 2006;25:7235-44. [Crossref] [PubMed]


