miR-125b prevent the progression of esophageal squamous cell carcinoma through the p38-MAPK signaling pathway
Introduction
Esophageal cancer (EC) is a highly prevalent digestive system cancer, increasing incidence, and mortality rates. Although the medical treatment and surgical techniques are developed too much in the last few years, the 5-year surviving percentage of EC patients is still low. One of the factors is that patients with EC have been diagnosed at an advanced stage or have localized invasive and distant metastases. EC can be separated into a couple of pathological types: esophageal squamous cell carcinoma (ESCC) and esophageal adenocarcinoma. The occurrence of ESCC is a multi-factor, multi-step biological process, often accompanied by the dormanting of tumor-inhibiting genes and oncogenes’ activation, like STAT3, PTEN, Snail1 et al. (1,2). The expression dysregulation of these genes greatly resembles the existence and progression of ESCC, but its pathogenesis is not clear.
With the development of genetics and molecular biology, people have a deeper understanding of the pathogenesis of ESCC. In recent years, microRNAs (miRNAs) have been implicated in tumorigenesis and tumor development. MicroRNAs are short non-coding RNAs, almost 18–25 nucleotides long, and are highly conserved (3). MiRNAs can regulate gene expression by the 3’UTR region of that particular mRNA, resulting in regulating translation suppression and/or mRNA degradation at the after-transcription stage, having relation with different life studies cell differentiation, development, and homeostasis process (4). Numerous studies have confirmed that miRNA plays a vital regulatory function in the development and progression of malignant tumors, and it resembles much to the diagnosis, treatment, and prognosis of tumor patients.
MiR-125b is examined in past ages to function as a tumor inhibitor in many cancers. In general, the outlook of miR-125b is down-regulated in tumors and plays a similar function as an oncogene, and the appearance of miR-125b is tissue specific. In different tumors comprising cells, the expression level of miR-125b is different (5), which makes miR-125b increasingly promising in the clinical testing and treatment and prognosis judgment of malignant tumors. At present, it has been found that miR-125b plays the role of oncogene or tumor suppressor gene in gastric cancer (6), liver cancer (7), colorectal cancer (8), breast cancer (9), lymphoma (10) and other malignant tumors, but its expression and regulatory role in ESCC are still unclear.
That is why this exhaustive study aimed to research the outlook of miR-125b in ESCC and its clinical benefits and study the effect of miR-125b on the biological function in ESCC development on a molecular level. The expression level of miR-125b in tumor comprising cells, some samples of ESCC was detected to analyze the interlink between miR-125b and the clinicopathological parameters and recovery as anticipated of patients with ESCC. Then cell experiments were conducted to explore the effect of miR-125b on the biological function of ESCC cells, and the potential target genes were predicted and verified by bioinformatics analysis.
We present the following article in accordance with the MDAR checklist (available at http://dx.doi.org/10.21037/jgo-20-546).
Methods
Cell line
Human ESCC KYSE410, EC-1, Eca109, TE-10, OE33 cells, and normal esophageal squamous epithelial human nonendometrioid carcinomas (NEECs) cells were bought from ATCC, USA. ESCC cells were nourished in DMEM medium holding fetal bovine serum almost 10% and added with 100 U/mL of penicillin along with 100 µg/mL of streptomycin. NEECs were cultured in RPMI medium containing fetal bovine serum almost 10% and placed in a CO2 incubator at 37 °C, 5% CO2, and cell culture medium kept changing after to 2 days of interval.
miR-125b‘s detection in cells by qRT-PCR
Total RNA was taken out from ESCC holding cells and ESCC cell lines utilizing Trizol regent, and 5µg of RNA was subjected to reverse transcription to obtain cDNA. The obtained cDNA was used as a template, and the RT-PCR reaction was conducted using SYBR Green PCR Kit. The reaction conditions around were 95 °C for 5 minutes; 40 cycles of approximately 95 °C, 30 seconds, 60 °C, 30 seconds, 72 °C, 1 minute. After the reaction was completed, the dissolution curve was drawn, and U6 was used as the internal reference. The expression of the miR-125b was then calculated by the 2−ΔΔCt method. The following primer pairs were used for the qPCR: miR-125b, forward: 5'-TCC AGT TCG GGA AGT GAA GTG A-3', and reverse: 5'-GTC CAA AGT GGT ATT GCC ACT A-3'. GAPDH, forward: 5'-CTC CAG TAC GTA AGT GCA GAG CT-3', and reverse: 5'-CAG TGA CAC ACG TTG CGT TGT C-3'.
Cell transfection
The cells were infused into all 6-well plates for 24 hours before transfecting it. When the cell fusion degree was 60% to 70%, the cells were then transfected, keeping in view the instructions given by Lipofectamine 2000 transfection reagent, and the cells were separated into four groups: miR-125b mimics, miR-125b mimics control, miR-125b inhibitor, and miR-125b inhibitor control. After transfecting it, the cells were cultured in a CO2 incubator at 37 °C, 5% CO2 for 48 hours for the next experiments. And RT-PCR was then used to detect transfection efficiency.
Cell proliferation assay
The transfected four groups of cells were made into cell suspensions, and after cell counting, 4×103 cells were seeded into plates, i.e., 96-well, and 3 of the replicate wells were set in every group. After the cells were all attached, the CCK-8 test kit was then used to detect the absorbance at the 0, 24, 48, and 72 hours of each cell‘s cell.
Transwell assay
Matrigel gel dissolved overnight at 4 °C was diluted 1:8 on a serum-free medium, and a Transwell chamber with an 8 µm microporous polycarbonate membrane was also put in a sterile plate of 24-well and added to the uppermost portion. The experimenter evenly spread 25 mol/L matrix gel on the bottom of the chamber, and sterile at 37 °C overnight to ensure sufficient polymerization of matrix gel. Transwell cell migration experiments were not performed with Matrigel gel. Each group of cells was collected and made into a single cell suspension. Every group of cells then was separately included in the uppermost portion of the Transwell chamber (1.5×105 cells/cell), and 650 µL of RPMI (medium) containing almost 10% of the fetal bovine serum was poured to the lower chamber. Forty-eight hours, the uppermost chamber cells were thoroughly cleaned utilizing cotton swab, washed with PBS, colored with crystal violet, examined using an inverted phase (contrast) microscope (×200), photographed, and five fields were randomly observed to count the number of transmembrane cells. The experiment was done three times.
Dual-luciferase report assay
A search of bioinformatics websites TargetScan and miRanda showed that miR-125b has a binding place for 3'URT of p38-MAPK, verified by dual luciferase assay, and the luciferase reporter gene was hybridized with wild-type p38-MAPK. The 3'UTR of the mutant p38-MAPK was combined, and the miR-125b was co-transfected with the plasmid carrying the p38-MAPK target fragment for 24 hours. Detection by a multi-function microplate reader and calculating the luciferase, firefly luciferase, and value/Renilla luciferase activities, value setting three duplicate wells per sample.
Western blotting
The total cell protein was extracted by RPPA cell supplementation with PMSF, and protein concentration was measured by the BCA method. The experimenter took 20 protein samples for SDS-PAGE electrophoresis, and the separated protein bands were transferred to PVDF membranes, blocked with Western blocking solution for 2 hours at room temperature; then rat anti-human p38-MAPK polyclonal diluted 1:1,000 was added. Antibody (1:1,000), the internal reference GAPDH antibody was diluted 1:1,000, and reacted at 4 °C overnight; then add 1:1,000 dilution of goat anti-mouse IgG (1:5,000), react 27–30 °C for 2 hours; BeyoECL chemiluminescence Detection reagent color development, chemiluminescence color imaging, and image analysis using Quantity One software.
Statistical analysis
All the collected information were presented as mean ± standard deviation (x±s) utilizing SPSS19.0 statistical software. Paired t-tests were then used for comparing the two groups. The difference coming between them was statistically better at P<0.05.
Results
miR-125b is shown in ESCC tissues and cell lines
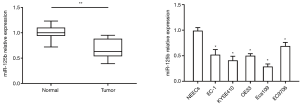
RT-PCR results proved that the relative appearance of miR-125b in ESCC tissues was significantly lower than that in adjoining normal tissues (P<0.05). Similarly, miR-125b was down-expressed in 5 ESCC cell lines equated with normal ESCC, statistically significant (Figure 1). It shows that miR-125b is downregulated in ESCC.
Inter relation analysis between miR-125b appearance and clinicopathological parameters
According to the median relative appearance of miR-124 in ESCC holding cells. We divided 52 patients between a very high expression category and also low expression category. The results concluded that the apparent position of miR-125b in tumor tissues was highly correlated between the strength of the invading of TNM level and lymph node metastasis, and no significant correlation with sex, age, and differentiation of tumor (Table 1).
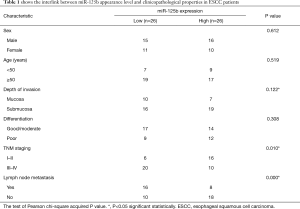
Full table
Interrelation analysis between miR-125b expression and patient recovery as expected
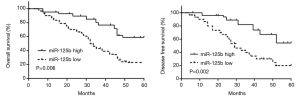
Kaplan-Meier curve analysis for existence and survival rate was used to see the link between miR-125b appearance and recovery as predicted of patients with ESCC. The results showed that ill patients with high miR-125b tend to have a longer survival period (Figure 2).
Cell proliferation
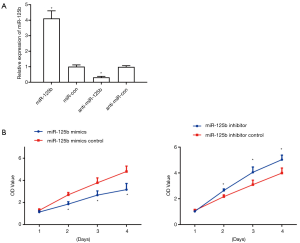
Eca109 cells were divided into four groups by cell transfection: miR-125b mimics, miR-125b mimics control, miR-125b binder, and miR-125b Clogger control, along with the transfection strength was verified by RT-PCR. The results concluded that the expression of miR-125b in the miR-125b mimics group was much more than the control group; on the other hand, the expression in the miR-125b Clogger group was much lesser than the control group, indicating successful transfection. The proliferation of transfected cells at the 0, 24, 48, and 72 hours was detected by the CCK8 method. The results concluded that miR-125b mimics cells‘ proliferation activity was much downsized from the 48th hour, while the proliferation activity of miR-125b Clogger group cells was much elevated (Figure 3), is showing that miR-125b can clog the proliferation of the ESCC cells.
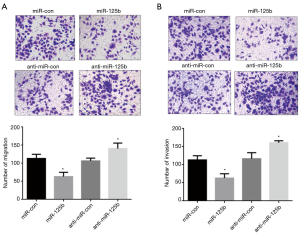
Transwell assay for cell migration and invasion
The Transwell cell migration assay outcome concluded that the number of transmembrane cells in the miR-125b mimics group was lesser than in miR-125b mimics control group transmembrane cells among the miR-125b clogger group was more than that in the miR-125b inhibitor control group (Figure 4A). Similarly, the Transwell cell invasion assay showed that after transfection of the miR-125b mimics, the percentage of Eca109 transmembrane cells in ESCC cells was downsized transmembrane cells transfected with miR-125b was much elevated (Figure 4B). It shows that miR-125b can inhibit the shifting and occupy of ESCC cells.
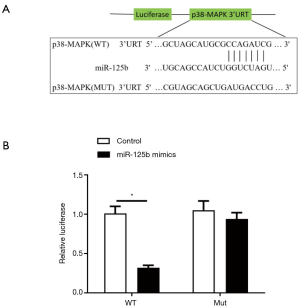
Dual-luciferase assay for miR-125b targeting p38-MAPK
Bioinformatics software analysis revealed a binding place for the miR-125b and p38-MAPK (Figure 5A). The outcome of dual-luciferase assay concluded that the relative fluorescence intensity of p38-MAPK in miR-125b mimics transfection group was lesser than in the control group (P<0.01), and the miR-125b binding site in p38-MAPK 3'UTR was also changed/mutated (Figure 5B). The clogging effect of the miR-125b over the reporter gene disappeared, showing that that miR-125b can act on 3'UTR of p38-MAPK messenger RNA directly to stop the expression of p38-MAPK.
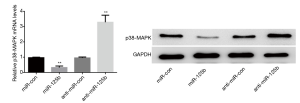
Outcome of the miR-125b on p38-MAPK mRNA and protein appearance
RT-PCR results concluded that the expression of p38-MAPK mRNA was downsized after more expression of miR-125b, while p38-MAPK mRNA level was raised after clogging of miR-125b expression, statistically significant (P<0.05); The outcomes of Western blotting indicated that the appearance of p38-MAPK protein in miR-125b mimics group was decreased, while the expression of the p38-MAPK protein in miR-125b inhibitor group had dropped significantly (P<0.05) (Figure 6).

Discussion
ESCC progression is a complex process involving multiple gene regulation and multiple steps. It is of huge clinical use to understand the mechanism of ESCC development and improve the initial stage diagnosis of ESCC and improve patients’ recovery expectations. MicroRNAs (miRNAs) are a category of small chain non-coding RNA molecules that are about 18–25 nucleotides inhibit length. By binding with the 3'URT site of a particular microRNA, and miRNAs cause mRNA degradation, inhibit protein translation, and regularizes gene showing after the transcriptional stage. Presently, several miRNAs have been found out, most of which are present in cells and have various biological functions such as regulating cell differentiation, development, and homeostasis. Studies found that most of the miRNAs are located in tumor-associated chromosomal sites, participate in regularizing transcriptional expression of tumor-associated genes, and thus participate in DNA damage of tumor cells, regulate the cell cycle, apoptosis and change the tumor microenvironment (11). In tumorigenesis and development, miRNAs play the role of oncogenes or tumor suppressor genes. Earlier studies have reported that more than 30% of human genes regulated with miRNAs aid (12), but the molecular process involved in regulating miRNAs is still largely unclear. MicroRNAs are tissue-specific, conserved, and time-series, making miRNAs have great clinical potential and can serve as potential markers for cancer diagnosis, therapeutic evaluation, and prognosis.
MiR-125b is a part of the miR-125 (family) residing in chromosome 21, which is affiliated with differentiation, proliferation, apoptosis, occupying, metastasis, and malignant cells’ drug resistance containing tumor (13). The expression of that miR-125b is tissue-specific, and miR-125b exerts different regulatory functions in different tumor tissues, such as bladder cancer, breast cancer, hepatocellular carcinoma, and ovarian cancer, melanoma, and osteosarcoma. And below-regulation of miR-125b exerts a tumor-suppressing effect, while in pancreatic cancer and prostate cancer, the expression of miR-125b is much higher than that of neighboring normal tissues and does a function in promoting cancer progression (14). Panella et al. found that the expression of miR-125b was regulated less in hepatocarcinoma containing cells and hepatocellular carcinoma cell lines, and the proliferation along with metastasis of hepatoma cells could be inhibited by targeting the oncogene lin28 (15). By overexpressing miR-125b in cancer cell SKBR3bof the breast, Vilquin et al. found that the appearance of ERBB2 and ERBB3 protein was decreased, and the proliferation ability of breast cancer cells was inhibited (16). Aherne et al. found that miR-125b was hijacked in prostate cancer cells. In androgen receptor prostate cancer cell lines LNCAP and CDs, androgen can develop the buildup of cancer cells of prostate by up-regulating miR-125b (17). The above studies reveal that miR-125b does indispensable regulatory work in developing various tumors, but its research in ESCC is still rare.
P38MAPK among the four major signaling pathways having a hand in MAPKs. Extracellular stimuli such as bacterial pathogens and cytokines activate the simultaneous phosphorylation of the Tyr and Tyr double sites of p38MAPK, and the activated p38MAPK transduces signals to Nuclei initiate transcription of related target genes and also transduce signals to other cellular components to produce different cellular responses (18). P38MAPK involves a huge scale biological process, i.e., promoting apoptosis, working in inflammatory reactions, ischemia-reperfusion injury, promoting cell phenotype transdifferentiation, and targeting a variety of different proteins (19). Activation of p38MAPK signaling pathway and manufacturing of pro-inflammatory cytokines [for instance interleukin (IL-1β), IL-6, Tumor Necrosis Factor-α (TNF-α)], induction of Cyclooxygenase 2 (COX-2), expression of nitric oxide synthase (infusible NOS) and intercellular adhesive molecule-1 (ICAM-1) is closely related (20). Recent studies have displayed that p38MAPK is highly expressed in breast cancer, prostate cancer, showing that it may be related to tumorigenesis and development. Chen et al. found that miR-3188 can target p38MAPK to inhibit outgrowth migration and invading breast cancer cells (21). Hu et al. found that activation of the p38MAPK signaling pathway in pancreatic cancer can increase tumor cells’ proliferative ability and promote peripheral blood vessels (22).
Here we first saw the expression percentage of miR-125b in ESCC tissues of the tumor. It was found that the level of expression of miR-125b in the tumor tissues was lesser than that in bordering normal tissues, and the decreased expression of miR-125b and the depth of tumor infiltration, TNM staging was relating to lymph node metastasis. The prognostic study showed that the survival period of miR-125b high expression was well higher than that of miR-125b low-level expression, indicating that miR-125b expression affects the degree of the continuity of disease and recover forecasted in patients with ESCC. Furthermore, the expression level of miR-125b in ESCC cell lines was determined by RT-PCR. The results gave the lesson that the expression of miR-125b was down-regulated in five ESCC cell lines, which was regular with the results detected in tumor tissues, indicating that miR-125b is down-regulated in ESCC. The expression of miR-125b in ESCC cell line Eca109 was successfully regulated by cell transfection. The proliferation, migration, and invasion ability of ESCC cells were discovered by the CCK-8 method and the Transwell method. The results showed that large-level expression of miR-125b, the accumulation/elevation, migration and entry, and invasion of ESCC cells decreased greatly, indicating that miR-125b can inhibit the proliferation, migration, and invasion of ESCC cells. MiR-125b plays a similar role as a tumor stopping gene. Bioinformatics predicts that p38-MAPK may be a targeting gene of miR-125b. Dual-luciferase results show that miR-125b has a focused targeting effect on p38-MAPK, and big scale expression of miR-125b can block p38-MAPK mRNA. And protein expression, showing that miR-125b has the quality to regulate the ESCC cells by p38-MAPK signaling pathway.
While summing up this discussion, the expression of miR-125b is down-regulated in the ESCC, and its expression strength is closely related to the depth of the invasion of the tumor, TNM level lymph node metastasis, and prognosis. miR-125b has this feature of stopping the proliferation, shifting, and invasion of ESCC cells by having a target p38-MAPK signaling pathway, which plays a similar role as a tumor suppressor gene.
Acknowledgments
Funding: None.
Footnote
Reporting Checklist: The authors have completed the MDAR checklist. Available at http://dx.doi.org/10.21037/jgo-20-546
Data Sharing Statement: Available at http://dx.doi.org/10.21037/jgo-20-546
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/jgo-20-546). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Chen MJ, Deng J, Chen C, et al. LncRNA H19 promotes epithelial mesenchymal transition and metastasis of esophageal cancer via STAT3/EZH2 axis. Int J Biochem Cell Biol 2019;113:27-36. [Crossref] [PubMed]
- Sherman SK, Maxwell JE, Qian Q, et al. Esophageal cancer in a family with hamartomatous tumors and germline PTEN frameshift and SMAD7 missense mutations. Cancer Genet 2015;208:41-6. [Crossref] [PubMed]
- Lau KW, Zeng H, Liang H, et al. Bioinformatics-based identification of differentiated expressed microRNA in esophageal squamous cell carcinoma. Transl Cancer Res 2018;7:1366-75. [Crossref]
- Rupaimoole R, Calin GA, Lopez-Berestein G, et al. miRNA deregulation in cancer cells and the tumor microenvironment. Cancer Discov 2016;6:235-46. [Crossref] [PubMed]
- Zuo D, Chen L, Liu X, et al. Combination of miR-125b and miR-27a enhances sensitivity and specificity of AFP-based diagnosis of hepatocellular carcinoma. Tumour Biol 2016;37:6539-49. [Crossref] [PubMed]
- Riquelme I, Tapia O, Leal P, et al. miR-101-2, miR-125b-2 and miR-451a act as potential tumor suppressors in gastric cancer through regulation of the PI3K/AKT/mTOR pathway. Cell Oncol (Dordr) 2016;39:23-33. [Crossref] [PubMed]
- Zhao L, Wang W. miR-125b suppresses the proliferation of hepatocellular carcinoma cells by targeting Sirtuin7. Int J Clin Exp Med 2015;8:18469-75. [PubMed]
- Tanaka H, Hazama S, Iida M, et al. miR-125b-1 and miR-378a are predictive biomarkers for the efficacy of vaccine treatment against colorectal cancer. Cancer Sci 2017;108:2229-38. [Crossref] [PubMed]
- Hong L, Pan F, Jiang H, et al. miR-125b inhibited epithelial-mesenchymal transition of triple-negative breast cancer by targeting MAP2K7. Onco Targets Ther 2016;9:2639-48. [PubMed]
- Zhan C, Wang T, You H, et al. Different expressions of miR-125b and SOX30 in malignant lymphomas and their significance. J BUON 2018;23:1179-84. [PubMed]
- Tang W, Wan S, Yang Z, et al. Tumor origin detection with tissue-specific miRNA and DNA methylation markers. Bioinformatics 2018;34:398-406. [Crossref] [PubMed]
- Yang L, Bian Y, Li Z, et al. Identification of potential biomarkers and pathways in ulcerative colitis with combined public mRNA and miRNA expression microarray data analysis. J Gastrointest Oncol 2019;10:847-58. [Crossref] [PubMed]
- Sun X, Li M, Sun Y, et al. The developmental transcriptome sequencing of bovine skeletal muscle reveals a long noncoding RNA, lncMD, promotes muscle differentiation by sponging miR-125b. Biochim Biophys Acta 2016;1863:2835-45. [Crossref] [PubMed]
- Zuberi M, Khan I, Mir R, et al. Utility of serum miR-125b as a diagnostic and prognostic indicator and its alliance with a panel of tumor suppressor genes in epithelial ovarian cancer. PLoS One 2016;11:e0153902. [Crossref] [PubMed]
- Panella M, Mosca N, Di Palo A, et al. Mutual suppression of miR-125a and Lin28b in human hepatocellular carcinoma cells. Biochem Biophys Res Commun 2018;500:824-7. [Crossref] [PubMed]
- Vilquin P, Donini CF, Villedieu M, et al. MicroRNA-125b upregulation confers aromatase inhibitor resistance and is a novel marker of poor prognosis in breast cancer. Breast Cancer Res 2015;17:13. [Crossref] [PubMed]
- Aherne ST, O’Neill F, Madden SF, et al. miR-125 family of miRNAs mediates prostate cancer cell proliferation and migration. Proceedings: AACR 107th Annual Meeting 2016.
- Tanaka T, Iino M, Goto K. Sec6 enhances cell migration and suppresses apoptosis by elevating the phosphorylation of p38 MAPK, MK2, and HSP27. Cell Signal 2018;49:1-16. [Crossref] [PubMed]
- Chen H, Wang X, Yan X, et al. LncRNA MALAT1 regulates sepsis-induced cardiac inflammation and dysfunction via interaction with miR-125b and p38 MAPK/NFκB. Int Immunopharmacol 2018;55:69-76. [Crossref] [PubMed]
- Xia P, Zhang R, Ge G. C/EBPβ Mediates TNF‐α‐Induced Cancer Cell Migration by Inducing MMP Expression Dependent on p38 MAPK. J Cell Biochem 2015;116:2766-77. [Crossref] [PubMed]
- Chen X, Chen J. MiR-3188 regulates cell proliferation, apoptosis, and migration in breast cancer by targeting TUSC5 and regulating the p38 MAPK signaling pathway. Oncol Res 2018;26:363-72. [Crossref] [PubMed]
- Hu H, Han T, Zhuo M, et al. Elevated COX-2 expression promotes angiogenesis through EGFR/p38-MAPK/Sp1-dependent signalling in pancreatic cancer. Sci Rep 2017;7:470. [Crossref] [PubMed]
(English Language Editor: J. Chapnick)