Cyclooxygenase-2 gene polymorphisms and susceptibility to colorectal cancer in a Brazilian population
Introduction
Colorectal cancer (CRC) is considered a public health problem worldwide (1). In Brazil, it was registered 15,067 death for this malignancy in 2013 (2) and the incidence is estimated at 34,280 people in 2016 (3).
The multi-ethnicity of Brazilian population represents a challenge in terms of genetic variability characterization, and the intra-individual genetic variation represented by genetic polymorphisms can be a relevant factor in susceptibility to developing cancer, distinct response to chemotherapy and radiotherapy, and prognosis (4). SNPs are the most common intra-species variants, exceeding 150 million cataloged in humans according to NCBI SNP database. SNPs may vary according to ethnic differences, explaining some discrepancies between the results found in several studies on this topic (5,6).
Cyclooxygenases are key enzymes of the inflammatory process, by converting arachidonic acid to prostaglandin H2, precursor of prostaglandins, prostacyclins and thromboxanes (7). The Cyclooxygenase-2 (COX-2), undetectable under normal conditions, is readily induced in response to growth and inflammatory factors, and it is expressed in inflammatory diseases, pre-malignant lesions and colorectal tumors (8). The enzyme may be detected in adenomas and adenocarcinomas (9). Recent studies have found COX-2 overexpression in up to 72% of CRC (10,11).
The promoter of the COX-2 gene is rich in recognition sites for nuclear proteins, which is critical in gene transcription induction (12). The COX-2-1195G > A (rs689466) SNP creates a recognition sequence for c-MYB transcription factor that results in increased gene transcription (12). This SNP has been associated to an increased risk of gastrointestinal tumors, including CRC (6,13).
The COX-2 + 8473T > C polymorphism (rs5275), can affect the affinity of transcription factor binding sites, influence the stability and/or translational efficiency of mRNA, and may modulate cancer susceptibility (14,15).
Thus, the aim of the present study was to evaluate the influence of COX-2 −1195A > G and 8473T > C polymorphisms as a risk factor of developing CRC.
Methods
Patients and controls
We evaluated 230 patients with CRC, who underwent surgical resection between September 2001 and November 2006, at the Hospital das Clinicas, University of São Paulo. The control group included 196 patients operated for benign disease at the same hospital, matched for sex and age, and no individual or familial history of cancer. All participants signed an informed consent form, approved by the Ethics Committee for Research Projects Analysis of the University of São Paulo, School of Medicine (protocol nº0803/11).
DNA isolation
Genomic DNA was isolated from the buffy coat using the extraction and purification Kit PureLink™ Genomic DNA Mini Kit (Invitrogen- Thermo Fisher Scientific, Carlsbad, USA) according to manufacturer’s instructions. The concentration and the purity of the DNA samples were determined in spectrophotometer NanoDrop™ ND-1000 (NanoDrop Technologies, Inc. Wilmington, USA). The integrity was checked by electrophoresis in 1% agarose gel.
Genotyping
Determination of genotype of COX-2 −1195A > G and 8473T > C SNPs was performed using TaqMan Kits (Thermo Fisher Scientific, Foster City, CA; Assay-on-demand, products: C_1647381_10 e C_16198794_10, respectively), according to manufacturer’s instructions. The selection of SNPs was based on association to CRC and functional effects described in the literature, available in the dbSNP database (NCBI; www.ncbi.nlm.nih.gov). Twenty percent of the samples were randomly selected and re-genotyped. The results showed 100% of similarity.
Statistical analysis
The genotype frequencies were determined by direct counting of the alleles. The Hardy-Weinberg equilibrium (HWE) was evaluated through χ2 test. The haplotype frequencies were calculated using the Expectation-Maximization (EM) algorithm. Linkage disequilibrium (LD) between polymorphisms was verified using the Haploview software (version 4.2). We evaluated the association of genotypes and alleles of the COX-2 gene between case and control groups using the χ2 test or Fisher’s exact test. The odds ratio and 95% confidence intervals were calculated for genotypes and estimated haplotypes. The logistic regression model adjusted for age, sex, ethnicity, level educational, alcohol drinking and smoking status were calculated in case and control groups. The genotype, dominant and recessive models were used in all analyzes, performed with R software, version 3.1.2. Data were considered significant when P<0.05.
Results
Demographics characteristics of the cases and controls are summarized in Table 1. There was no statistical difference between the groups regarding age (P=0.84), sex (P=1.00), ethnicity (P=0.58) and smoking status (P=1.00). Education level and alcohol consumption showed statistical difference (P<0.001).
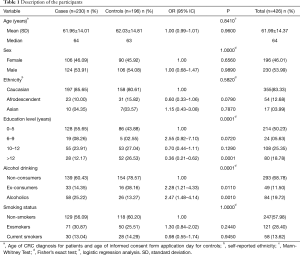
Full table
The position of the COX-2 polymorphisms on chromosome, information about HWE and minor allele frequency are shown in Table 2. The genotype frequencies of the COX-2 SNPs were consistent with HWE with P-value ≥0.05, except for control group of the 8473T > C SNP (P=0.02).

Full table
We also evaluated the allelic segregation of the polymorphisms, in order to verify the LD. There is a high LD between both polymorphisms, which are distant from each other in 7 kilobase (D’ =97). The two SNPs were grouped into a single block and formed three haplotypes, as shown in Figure 1.
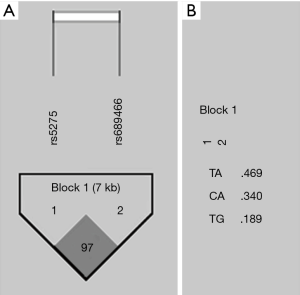
We determined the genotype frequencies of the CRC patients and control subjects. The results are summarized in Table 3. They were similar in both logistic regression models (P>0.05). The unadjusted model showed a trend to lower risk, but without statistical significance, for the 8473TT genotype (P=0.05).
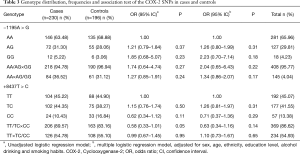
Full table
We evaluated the association between alleles of the COX-2 SNPs in cases and controls. No statistically significant differences were detected among groups (P>0.05). The same results were obtained with the haplotypes in both simple and multiple logistic regression models, as shown in Table 4.
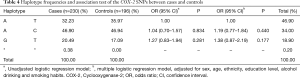
Full table
Discussion
This is the first report of the impact of COX-2 −1195A > G and 8473T > C polymorphisms in CRC risk among Brazilian population.
In the present study, the frequencies of the COX-2 SNPs were consistent with HWE in the general population, except for the SNP 8473T > C in control group (P=0.02). Deviation of HWE may be caused by interbreeding, population stratification, natural selection or unknown errors of the genotype determination among others (16). The Brazilian population may have poor compatibility for genetic ancestry among study groups because it is originated from a combination of many ethnic groups (17). Nonetheless, there are no standard guidelines for rejecting SNPs that deviate from HWE (17).
We evaluated the association between alleles of the COX-2 SNPs in cases and controls, as well as genotypes and haplotypes, and there was no statistically significant association to CRC risk. Functional studies have demonstrated the effect of the SNPs in major loci in the COX-2 gene, and these may be related to increased levels of mRNA in CRC (18). The SNP −1195A > G lies in a region rich in recognition sites for nuclear transcription factors, that are critical in the transcription activation (12).
Our results are similar to others findings identified in the literature. Hoff et al. (19) found no significant association of the COX-2 −1195A > G SNP with the risk of CRC in the Dutch population. Similar results were found in the Spanish and Danish population (20,21) and in two systematic reviews (22,23). In contrast, some researches detected significant association, but the results are conflicting (5,6). A case-control study in Jordan associated protective effect on the development of polyps and CRC in the presence of wild-type allele −1195A (24). Pereira et al. associated the G allele to predisposition to CRC increased in 1.73 fold (25). These results corroborate the findings of a functional study that identified overexpression of COX-2 gene in cell lines carrying the G allele (26). However, Tan et al. (13) associated the AA and GA genotypes with increased predisposition of CRC (OR =1.77 and 1.24, respectively), and the same association was detected in a systematic review (6). Vogel et al., showed that carriers of the G allele had a reduced risk of CRC (18). The allele creates a site for the transcription factor c-MYB, which would cause the increase of transcripts according to Zhang et al.(12).
The 8473T > C SNP may modify the binding affinity for regulatory factors and influence the mRNA stability and/or translational efficiency (14). This polymorphism was not associated to susceptibility to CRC in this study. Cox et al. (27) evaluated COX-2 SNPs, including the 8473T > C, and did not detect association. The same results were observed in four European studies (18,20,21,25) and in two systematic reviews (22,23).
The divergent results in different population and tumor types suggest that the SNPs play a role in CRC, but are influenced by other molecular factors including functional polymorphisms not yet described, which can increase or neutralize the function effect of the SNPs COX-2 −1195A > G and 8473T > C (12). Similarly there may be a cell-tissue-specific modulation, which could explain the different findings related to mechanism of action of the variants in gene and protein expression (28). Another possible explanation may be associated to interference of environmental factors and ethnic composition of the population, demonstrated by studies results in different countries and ethnic groups (29).
This study has some limitations. The number of cases was relatively small for this study design, which may result in decreased statistical power. SNPs were selected according to their frequency in Caucasian populations with more than 10% of polymorphic frequency to minimize the small sample size. Moreover, ethnicity was self-reported by the patient. The most appropriate method to determine this variable is through the characterization of informative autosomal markers of ancestry, which brings more specific information, especially in populations with a high degree of admixture, as in Brazil.
Therefore, under the conditions of this research, we concluded that the variants COX-2 −1195A > G and 8473T > C do not participate in the genetic susceptibility to CRC in the Brazilian population.
Acknowledgements
Funding: Major support for this study was provided by São Paulo Research Foundation (FAPESP), grant #2011/23181-5.
Footnote
Conflicts of Interest: The authors have no conflicts of interest to declare.
Ethical Statement: All participants signed an informed consent form, approved by the Ethics Committee for Research Projects Analysis of the University of São Paulo, School of Medicine (protocol nº0803/11).
References
- Ferlay J, Soerjomataram I, Ervik M, et al. GLOBOCAN 2012 v1.0, Cancer Incidence and Mortality Worldwide: IARC CancerBase No. 11, International Agency for Research on Cancer, Lyon, France, 2013.
- Brasil. Instituto Nacional do Cancer José Alencar Gomes da Silva. Estimativa 2016: Incidência de Câncer no Brasil. Available online: http://www.inca.gov.br/estimativa/2016/sintese-de-resultados-comentarios.asp
- Brasil, Ministério da Saúde, Instituto Nacional de Câncer. Atlas de mortalidade por cancer, 2013. Available online: http://www1.inca.gov.br/vigilancia/mortalidade.html
- Shields PG, Harris CC. Molecular epidemiology and the genetics of environmental cancer. JAMA 1991;266:681-7. [Crossref] [PubMed]
- Pereira C, Medeiros RM, Dinis-Ribeiro MJ. Cyclooxygenase polymorphisms in gastric and colorectal carcinogenesis: are conclusive results available? Eur J Gastroenterol Hepatol 2009;21:76-91. [Crossref] [PubMed]
- Dong J, Dai J, Zhang M, et al. Potentially functional COX-2-1195G>A polymorphism increases the risk of digestive system cancers: a meta-analysis. J Gastroenterol Hepatol 2010;25:1042-50. [Crossref] [PubMed]
- Vane JR, Bakhle YS, Botting RM. Cyclooxygenases 1 and 2. Annu Rev Pharmacol Toxicol 1998;38:97-120. [Crossref] [PubMed]
- Wang D, Mann JR, DuBois RN. The role of prostaglandins and other eicosanoids in the gastrointestinal tract. Gastroenterology 2005;128:1445-61. [Crossref] [PubMed]
- Balbinotti RA, Ribeiro U, Sakai P, et al. hMLH1, hMSH2 and cyclooxygenase-2 (COX-2) in sporadic colorectal polyps. Anticancer Res 2007;27:4465-71. [PubMed]
- Kim NK, Park JK, Shin E, et al. The combination of nuclear factor kappa B, cyclo-oxygenase-2 and vascular endothelial growth factor expression predicts poor prognosis in stage II and III colorectal cancer. Anticancer Res 2014;34:6451-7. [PubMed]
- Soumaoro LT, Uetake H, Takagi Y, et al. Coexpression of VEGF-C and Cox-2 in human colorectal cancer and its association with lymph node metastasis. Dis Colon Rectum 2006;49:392-8. [Crossref] [PubMed]
- Zhang X, Miao X, Tan W, et al. Identification of functional genetic variants in cyclooxygenase-2 and their association with risk of esophageal cancer. Gastroenterology 2005;129:565-76. [PubMed]
- Tan W, Wu J, Zhang X, et al. Associations of functional polymorphisms in cyclooxygenase-2 and platelet 12-lipoxygenase with risk of occurrence and advanced disease status of colorectal cancer. Carcinogenesis 2007;28:1197-201. [Crossref] [PubMed]
- Dixon DA, Kaplan CD, McIntyre TM, et al. Post-transcriptional control of cyclooxygenase-2 gene expression. The role of the 3'-untranslated region. J Biol Chem 2000;275:11750-7. [Crossref] [PubMed]
- Moore AE, Young LE, Dixon DA. A common single-nucleotide polymorphism in cyclooxygenase-2 disrupts microRNA-mediated regulation. Oncogene 2012;31:1592-8. [Crossref] [PubMed]
- Xu J, Turner A, Little J, et al. Positive results in association studies are associated with departure from Hardy-Weinberg equilibrium: hint for genotyping error? Hum Genet 2002;111:573-4. [Crossref] [PubMed]
- Lewis CM, Knight J. Introduction to genetic association studies. Cold Spring Harb Protoc 2012;2012:297-306. [Crossref] [PubMed]
- Vogel LK, Sæbø M, Høyer H, et al. Intestinal PTGS2 mRNA levels, PTGS2 gene polymorphisms, and colorectal carcinogenesis. PLoS One 2014;9:e105254. [Crossref] [PubMed]
- Hoff JH, te Morsche RH, Roelofs HM, et al. COX-2 polymorphisms -765G-->C and -1195A-->G and colorectal cancer risk. World J Gastroenterol 2009;15:4561-5. [Crossref] [PubMed]
- Siezen CL, Bueno-de-Mesquita HB, Peeters PH, et al. Polymorphisms in the genes involved in the arachidonic acid-pathway, fish consumption and the risk of colorectal cancer. Int J Cancer 2006;119:297-303. [Crossref] [PubMed]
- Andersen V, Ostergaard M, Christensen J, et al. Polymorphisms in the xenobiotic transporter Multidrug Resistance 1 (MDR1) and interaction with meat intake in relation to risk of colorectal cancer in a Danish prospective case-cohort study. BMC Cancer 2009;9:407. [Crossref] [PubMed]
- Wang J, Guo X, Zhang J, et al. Cyclooxygenase-2 polymorphisms and susceptibility to colorectal cancer: a meta-analysis. Yonsei Med J 2013;54:1353-61. [Crossref] [PubMed]
- Peng Q, Yang S, Lao X, et al. Meta-analysis of the association between COX-2 polymorphisms and risk of colorectal cancer based on case-control studies. PLoS One 2014;9:e94790. [Crossref] [PubMed]
- Shomaf M, Yousef AL, Ababna N, et al. Cyclooxygenase-2 (COX2) gene polymorphisms and the risk of sporadic colorectal cancer and polyps among Jordanian population. Turk J Gastroenterol 2015;26:154-8. [Crossref] [PubMed]
- Pereira C, Queirós S, Galaghar A, et al. Genetic variability in key genes in prostaglandin E2 pathway (COX-2, HPGD, ABCC4 and SLCO2A1) and their involvement in colorectal cancer development. PLoS One 2014;9:e92000. [Crossref] [PubMed]
- Pereira C, Sousa H, Silva J, et al. The -1195G allele increases the transcriptional activity of cyclooxygenase-2 gene (COX-2) in colon cancer cell lines. Mol Carcinog 2014;53 Suppl 1:E92-5. [Crossref] [PubMed]
- Cox DG, Pontes C, Guino E, et al. Polymorphisms in prostaglandin synthase 2/cyclooxygenase 2 (PTGS2/COX2) and risk of colorectal cancer. Br J Cancer 2004;91:339-43. [PubMed]
- Szczeklik W, Sanak M, Szczeklik A. Functional effects and gender association of COX-2 gene polymorphism G-765C in bronchial asthma. J Allergy Clin Immunol 2004;114:248-53. [Crossref] [PubMed]
- Sansbury LB, Millikan RC, Schroeder JC, et al. COX-2 polymorphism, use of nonsteroidal anti-inflammatory drugs, and risk of colon cancer in African Americans (United States). Cancer Causes Control 2006;17:257-66. [Crossref] [PubMed]


